Large scale simulation workflows are among the primary applications running on modern High-Performance Computing (HPC) facilities. Traditionally, HPC centres offer a batched execution model orchestrated by a central queue manager, which is not compatible with the human-in-the-loop pattern required by interactive notebooks.
Still, the possibility to monitor and interact with a running simulation can be crucial. For instance, a user can adjust diverging workloads at runtime to recover a correct state or early-stop them to avoid wasting time and resources. In addition, a simulation can be refined for better exploring a specific portion of the solution space.
A solution adopted by some HPC facilities worldwide is to install a Jupyter-based service (e.g. a JupyterHub instance) on the login nodes of the data centre or some publicly exposed instances of a tightly coupled Cloud service. Appropriate authentication chains percolate the user identity from the web interface to some HPC worker nodes made available on-demand by the queue manager. These modifications to the data centre control plane are non-trivial, mainly due to their security implications, and require the intervention of system administrators.
Jupyter Workflow comes with a series of advantages:
- Each code cell can be repeatedly offloaded to a remote HPC facility for large scale executions;
- Users can rely on the high-level visualisation tools provided by Jupyter Notebooks to examine the results interactively;
- Multiple versions of the same cell can be processed in parallel with different parameters through the Scatter/Gather pattern.
The Quantum ESPRESSO suite
In this article, a Quantum ESPRESSO simulation workflow is used to represent a broad class of traditional HPC Molecular Dynamics Simulation (MDS) tools. Quantum ESPRESSO is an integrated suite of Open-Source computer codes for electronic-structure calculations and materials modelling at the nanoscale. Quantum ESPRESSO has been shown to scale well to petascale systems, and it is currently addressing the exascale challenge.
As for other scientific libraries, much of the Quantum ESPRESSO performance derives from the linked matrix multiplication libraries (e.g. BLAS and LAPACK), which are tightly coupled with the underlying hardware technology at compile time. Tweaking performances of these libraries is out of reach of many domain experts, but linking Quantum ESPRESSO with low-performing or badly-compiled versions can have a massive impact on the time-to-solution. Offloading computation to an already optimised execution environment is a way to free users from this burden.
The primordial soup pipeline
To assess the capabilities of Jupyter Workflow to enable interactive simulations of realistic, large-scale systems, we chose a simulation of the so-called “primordial soup”. In particular, the workflow implements a Car-Parrinello simulation of a mixture of H2O, NH3 and CH4 molecules to represent the basic ingredients of life.
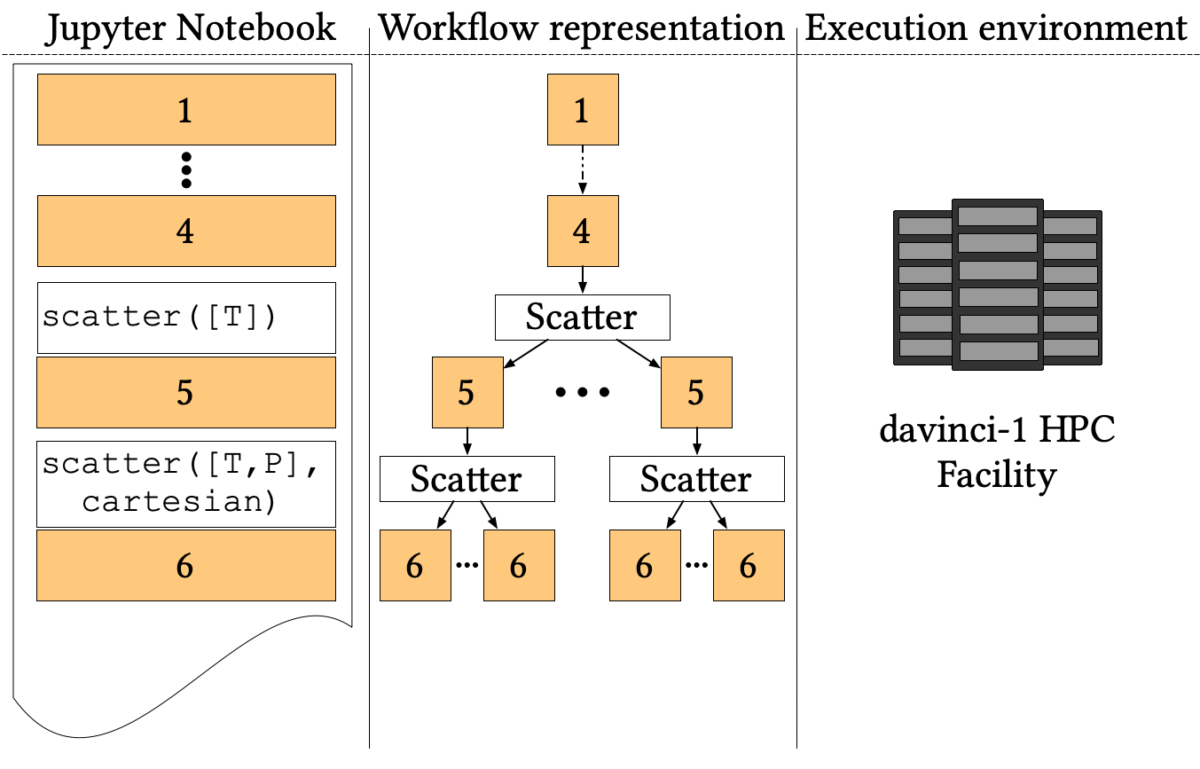
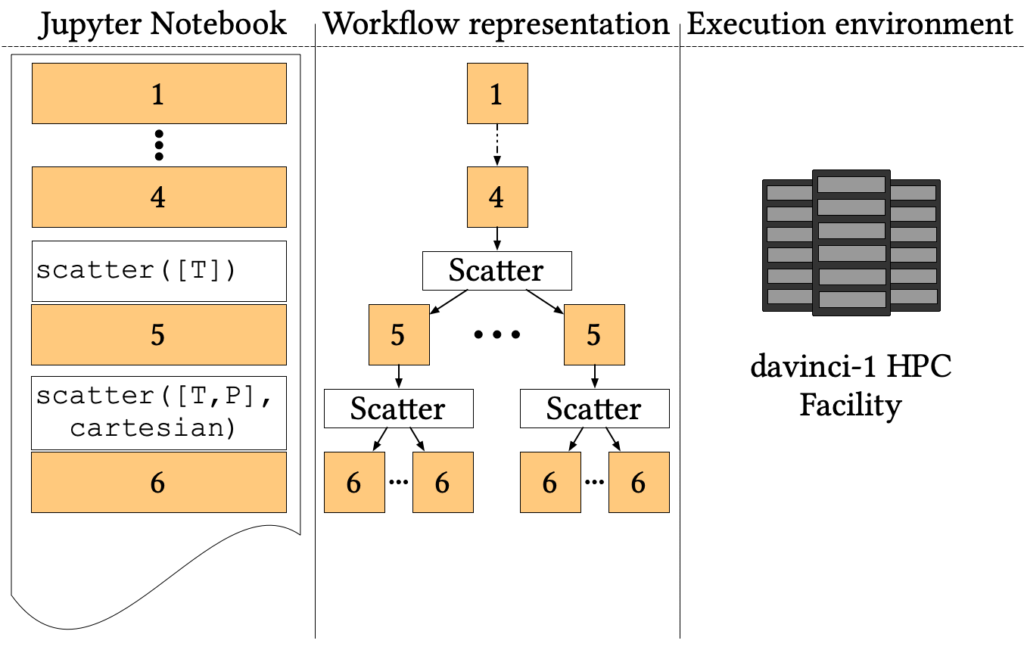
The simulation aims to explore the phase space to find where C-H, O-H and N-H bounds break up, forming more complex organic molecules. Several Car-Parrinello simulations at different pressure-temperature points (P, T) are needed to simulate the phase diagram. The workflow proceeds as follows:
- The first four cells, common to all simulations, prepare a starting state at room temperature and pressure from a random distribution of three molecules;
- Then the pipeline forks to simulate different temperatures through Nosé-Hoover thermostats (step 5);
- Finally, for each temperature, the simulation forks again to simulate each temperature at several values of pressure using the Parrinello-Rahman constant pressure Lagrangian (step 6).
The Notebook implementing this pipeline is available online on the official Jupyter Workflow GitHub repository.
Jupyter Workflow execution
We offloaded the execution of each step to two CPU nodes of davinci-1, the Leonardo S.p.A. HPC system. Each node is equipped with 2 Intel Xeon Platinum 8260 sockets (24 cores, 2.40 GHz each) and 1 TB of RAM. Since davinci-1 is not reachable from outside the company private network, we ran the Jupyter Workflow kernel on one of the login nodes.
We analysed the weak scalability of the application by running it on 1, 4 and 16 (P, T) points, comparing for each setting the time to complete steps 5 and 6 with the bare queue manager of the system, with interactive execution and with the optimised bulk evaluation offered by Jupyter Workflow.
The overhead introduced by the interactive execution mode becomes predominant with 16 (P, T) points, because data movements between the Jupyter Workflow control plane and the HPC file system are not negligible. Still, the overhead remains totally negligible with the bulk evaluation mode.
I. Colonnelli, M. Aldinucci, B. Cantalupo, L. Padovani, S. Rabellino, C. Spampinato, R. Morelli, R. Di Carlo, N. Magini and C. Cavazzoni, “Distributed workflows with Jupyter”, Future Generation Computer Systems, vol. 128, pp. 282-298, 2022. doi: 10.1016/j.future.2021.10.007